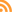- Author: Michael D Cahn
Introduction
Traditional winter cereal cover crops planted in the Salinas valley have many potential benefits including, scavenging nitrate in the soil profile, increasing organic matter in the soil, and protecting the soil from erosion during storm events. However, when grown for 3 to 4 months during the late fall and winter, cereal rye, triticale, or barley can accumulate 5 to 6 tons of dry matter biomass that must be incorporated into the soil before planting a spring vegetable crop. Tilling in a high amount of cover crop biomass can be disruptive to spring planting schedules. Consequently, only a small fraction of the vegetable ground in the Salinas valley is cover cropped each year.
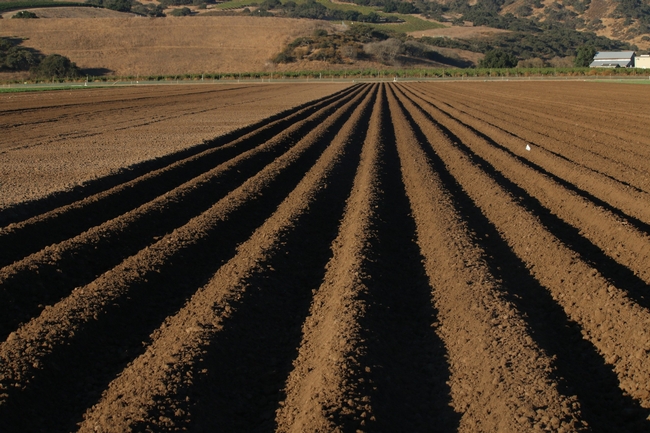
Previous studies demonstrated alternative strategies can limit the biomass growth of these cereal cover crop species so that they can more easily be tilled into the soil, and therefore less disruptive to spring planting schedules. After fall land preparation, the cereal cover crops are seeded into listed beds and/or in the furrow bottoms. After they become established they can reduce runoff and protect the soil from erosion during early winter storm events. Before the cover crops grow too big, they are terminated with an herbicide to limit the amount of above ground biomass that needs to be incorporated in the spring. For organic systems, planting a mustard cover crop on listed beds or furrows which can be terminated mechanically by mowing is another strategy to limit biomass. A good target for these low biomass cover crops is between 0.5 to 1 ton of dry matter per acre by the date of termination. Once terminated, the biomass begins to decompose. However, the residue on the surface continues to protect the soil from erosion and can significantly increase infiltration from rain events. This helps to leach accumulated salts in the soil as well as recharge groundwater aquifers. The remaining decomposed residue can easily be incorporated into the soil during bed preparation in the spring.
One risk of this low biomass approach is accessing fields during the winter to terminate the cover crop. If soil conditions are too wet or if there is not enough available labor, it may be difficult to fit in a spray application or to run a flail mower. This termination step also increases the cost of managing the cover crop. A possible solution is to use species that grow slowly during the winter when temperatures are cold. Sudangrass and sorghum-sudangrass hybrid are warm season adapted species that could be used in this low biomass approach to managing winter cover crops.
Field trial with warm season adapted cover crop species
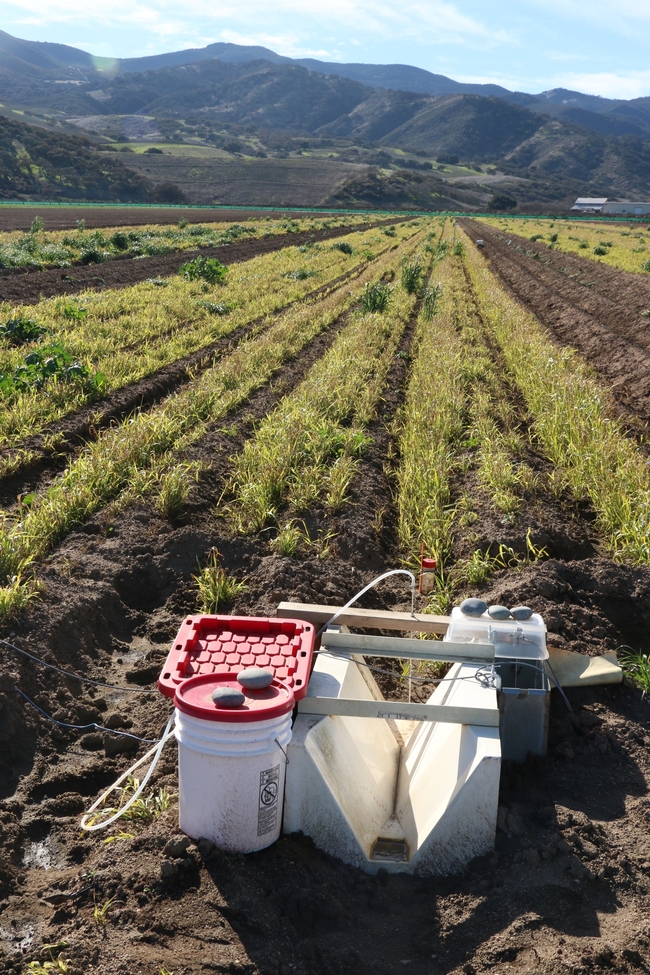
A field trial was conducted with sudangrass and sorghum-sudangrass in the 2023-2024 winter to evaluate biomass growth, and the effect on storm water runoff and soil erosion compared to bare-fallow plots. The site was located on an Arroyo Seco gravelly loam soil with a slope of more than 5%. Plots measuring 1050 ft in length by four 40-inch wide peaked beds were planted with either sudangrass, sorghum sudangrass hybrid, or left bare fallow. Treatments were replicated 4 times. The cover crops were seeded at 60 to 80 lbs/acre on October 4th and were subsequently sprinkle-irrigated several times. Total water applied for establishment was 2.6 inches. One application of the herbicide Bromoxnil (Maestro) was applied about 45 days after planting to kill emerged broadleaf weeds.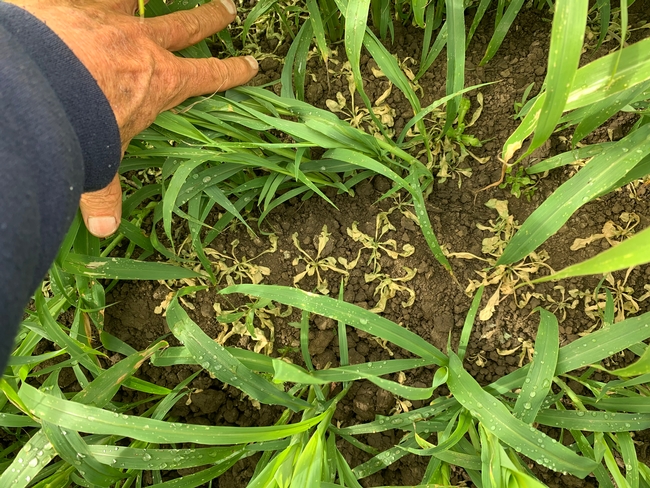 
Results
Above ground biomass, N uptake, and carbon accumulation
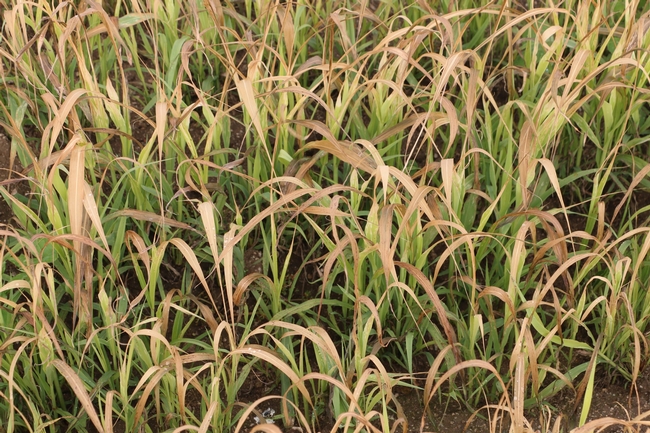
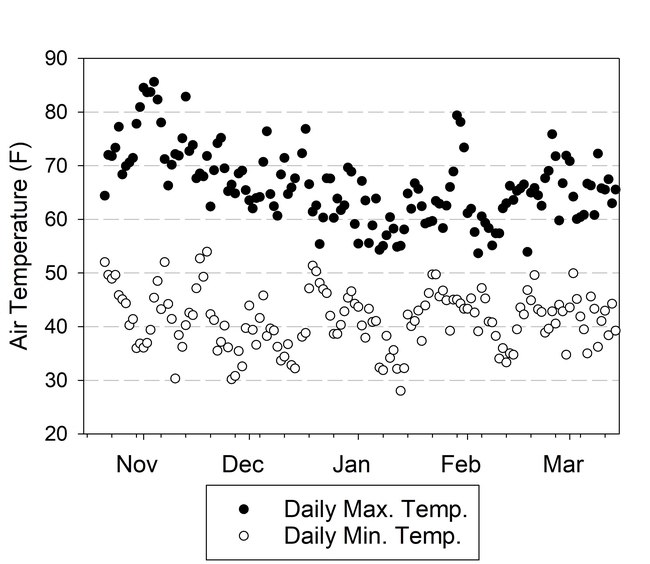
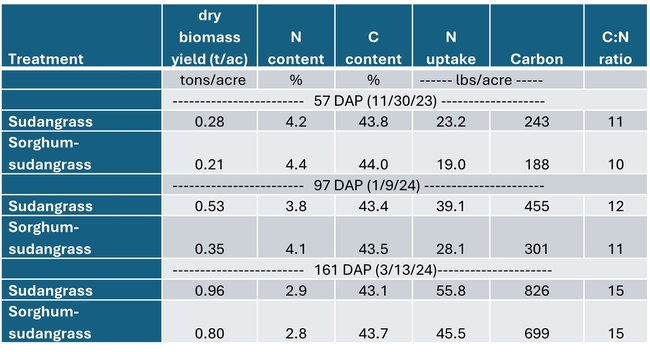
Both cover crops had limited biomass growth, accumulating only 0.35 to 0.5 tons/acre of dry matter by early January and less than 1 ton/acre by mid March (Table 1). Growth was set back by cold conditions that occurred from mid November through early January, occasionally reaching freezing temperatures which caused damage to leaves (Fig. 1). However, the freezing temperatures lasted only a few hours and were not severe enough to kill the cover crops (Fig. 2). By March 13th the cover crops had taken up 45 to 55 lbs N/acre and had a carbon to nitrogen ratio of 15. The C:N ratio of 15 would suggest that after soil incorporation the residue would decompose rather quickly and release N for the following vegetable crop.
Runoff, rainfall infiltration, and control of soil erosion
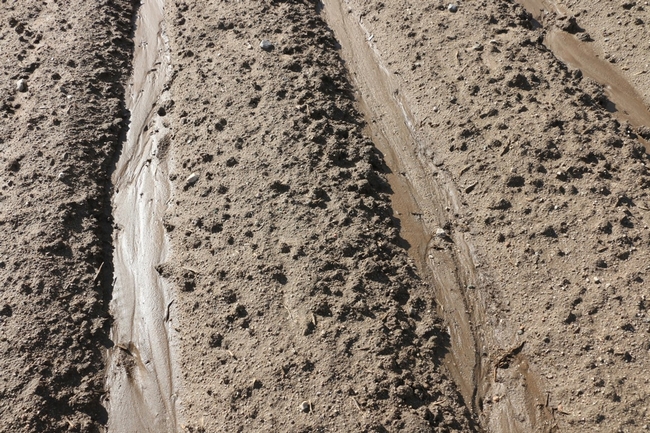
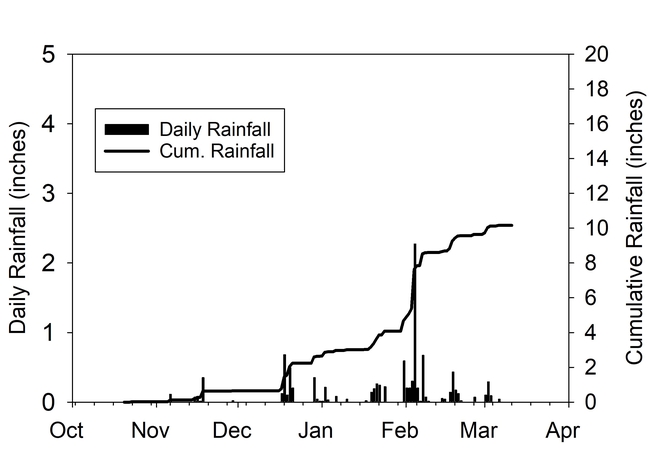
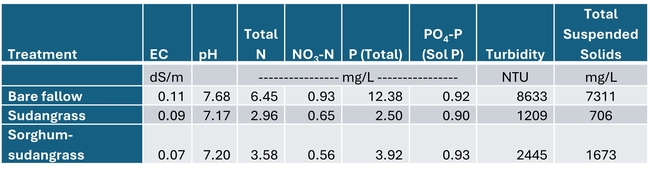
Total rainfall measured at the trial site was 10.2 inches for the winter season. The most intense period of rainfall occurred in late January and early February which resulted in several runoff events (Fig. 3). During this period about 50% of the rainfall in the bare fallow plots was lost as runoff compared to 15% lost as runoff in the cover crop plots (Fig.4). Over the entire winter season, runoff was reduced by an average of 70% under the cover cropped plots compared to the bare fallow plots, and significantly more rainfall was infiltrated into the ground in the cover cropped plots. In addition, suspended sediment concentration was 90% and 77% less in the sudangrass and sorghum-sudangrass cover crop plots, respectively, compared to the bare plots. Turbidity, total P, and total N concentration in the runoff were also reduced under the cover crop plots compared to the bare fallow plots (Table 2).
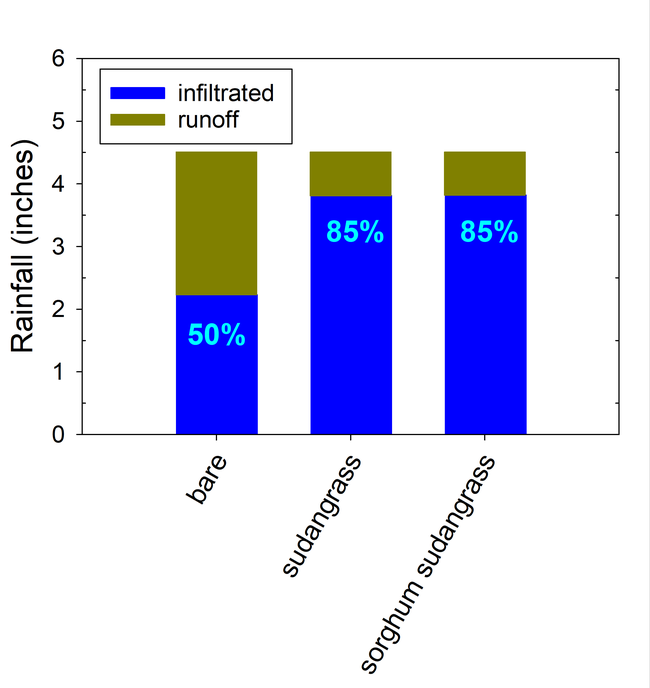
Seasonal soil erosion losses could be calculated based on the volume of the runoff and sediment concentration in the runoff. The total loss of sediment averaged more than 3500 lbs per acre in the bare fallow plots during the winter, while erosion losses were reduced by 96% to 98% in the sorghum-sudangrass and sudangrass plots (Fig. 5). Total N losses were reduced by 83% to 86% in the cover crop plots compared to the fallow plots, and total P losses were reduce by 81% to 85% in the cover cropped plots compared to the bare fallow plots.
Conclusions
The use of warm season species such as sudangrass and sorghum-sudangrass hybrids as winter cover crops provides several advantages compared to planting cereal cover crops. The biomass growth through the winter is self-limiting due to the cold conditions that typically occur in the Salinas Valley. Because the final biomass would likely be less than 1 ton per acre, these species can be planted on listed beds in the fall rather than on flat ground. This means that in the spring, the remaining cover crop can be lillistoned into the peaked beds a few weeks before final bedshaping and planting. Cover crops planted on flat ground and have high amounts of biomass usually require many tillage passes to prepare ground for planting. Despite, having less biomass than traditional winter cereal species, sudangrass and sorghum-sudangrass hybrid cover crops provided excellent erosion control compared to leaving the ground bare, and increased infiltration of rainfall during storm events. Also these species may be able to scavenge significant amounts of nitrogen from the soil which can limit nitrate leaching during the winter months.
On the east-side of the Salinas Valley groundwater levels have been in the decline for several decades. Infiltrating as much rainfall as possible during the winter using strategies such as low biomass cover crops could potentially help recharge the aquifer in this region. We plan to conduct a second year of field trials with these warm season species to continue evaluating this approach to managing winter cover crops in vegetable systems.
Acknowledgments
This project was funded by the California Leafy Greens Research Board.
- Author: Dylan Beal
Hello everyone, I'm the new UC IPM Entomology Advisor serving the communities of Monterey, Santa Cruz, and San Benito Counties. Born and raised in the Santa Clarita Valley, I'm excited to return to my home state of California.
Inspired by a lifelong passion for insects, I pursued bachelor of science in degrees in Entomology and Plant Pathology at Cornell University in Ithaca, NY. At Cornell, I was first exposed to the world of cooperative extension, working in a lab to study and support natural enemies of spider mites, aphids, thrips, fungus gnats, and whiteflies in greenhouses.
In partnership with Montpellier SupAgro and the Università degli Studi di Torino, I received a Master of Science degree in Viticulture and Enology. For my master's thesis, I worked with Italian grape growers on understanding the ecology and management of leafhopper vectors of the pathogen responsible for European grapevine yellows (Flavescence dorée phytoplasma).
For my PhD, I studied Environmental Science, Policy, and Management at the University of California, Berkeley. At Berkeley, I worked with grape growers and cooperative extension advisors in Napa and Sonoma county to understand the role of spittlebug insect vectors in spreading Xylella fastidiosa, the bacteria responsible for Pierce's disease of grapevine.
After Berkeley, I have worked as a postdoc at Washington State University with cherry and blueberry growers to adapt and adopt innovative and internationally recognized practices for sustainable management of Spotted-Wing Drosophila (SWD). As part of that work, I have closely partnered with growers, pest control advisors, USDA scientists, cooperative extension advisors, and industry representatives on SWD projects related to biological control, cultural control, behavioral control, and pesticide resistance screening and management. Additionally, I was part of a team that worked to coordinate SWD management with the management of the leafhopper vectors of X-Disease phytoplasma in cherry.
With 15 years of experience in New York, California, Washington State, France, and Italy, I'm excited to collaborate with you all to support the development and adoption of innovative, cost-effective, and sustainable insect pest management practices. I aim to develop partnerships, lifelong relationships, and projects that center and elevate grower voices, experiences, and leadership. Together, I hope that we can not only address the threat that agricultural insect pests pose to our three counties but also help to support the many communities and ecosystems that interface with our agricultural livelihoods. Please feel free to reach out to me at 831-759-7359 or djbeal@ucanr.edu. The Bug Doctor is in to look at any insects you may find!
- Author: Paramveer Singh
- Author: Richard Smith
University of California Cooperative Extension
2024 Automated Technology Field Day
Thursday, June 27, 2024 9:00 a.m. to 12:00 noon
Hartnell East Campus, Salinas, CA
1752 East Alisal Road, Salinas (follow signs to demonstration area)
Topics
- Growing role of automated technologies in precision weeding and other ag operations
- Automated weeders, thinners and sprayers from several companies will be demonstrated on planted crops
- Technologies appropriate for conventional and organic production will be demonstrated
- There will be ample opportunity to watch demonstrations of each machine and to discuss with company representatives.
Companies represented
- Boden Bolt
- Carbon Robotics
- Ecorobotix (Keithly Williams)
- L&Aser
- Mantis Ag Technology
- Smart Steam Applicator Organizers
- Sutton Ag
- Stout Industrial Technology
- Tensorfield
- Verdant Robotics
- Veda Farming
- Weed Spider
Organizers
- Steve Fennimore, Extension Vegetable Weed Control Specialist, UC Davis.
- Paramveer Singh, Agricultural Technology Area Advisor, Salinas.
- In cooperation with the USDA ARS, Salinas.
CA 3.0 Continuing Education Credits have been requested.
No registration fee and refreshments will be served.
For more information, call Steve Fennimore at (831) 594-1333.
2024 Automated Tech Field Day
- Author: Michael D Cahn
Date: Wed. April 3rd, 2024
Course 8:30 am – 12 pm
Location: Monterey County Agriculture Conference Room
1432 Abbott St, Salinas CA 93901
- Learn how to use CropManage to support irrigation and nutrient management decisions and record-keeping for your crops.
- Learn how to set up your ranch and crops in CropManage.
- Learn about the latest updates and how CropManage can assist with Ag Order 4.0
CropManage is a free online decision-support tool for water and nutrient management of vegetables, berry, agronomic, vineyard and tree crops. Based on research and field studies conducted by the University of California Cooperative Extension, CropManage provides real-time recommendations for efficient and timely irrigation and fertilization applications while maintaining or improving overall yield.
At this free workshop, we will provide hands-on training so that you can learn to use the newest version of CropManage. Crops currently supported include many vegetables (carrots, cabbage, celery, broccoli, lettuce, tomato, spinach, etc.), berry crops (raspberry and strawberry), vineyards, tree crops (almond, walnut, pistachio, prunes, and pear), and agronomic crops (alfalfa and corn). CropManage is also available in Spanish.
Who should participate? Growers, farm managers, other farm staff, crop advisors, consultants, and technical service providers are welcome. The workshop is for both new and current CropManage users. Spanish translation will be available.
What to bring? This is a participatory workshop. Please bring a tablet or laptop computer so that you can follow along and participate in the exercises. Each participant will need a user account for CropManage. Please set up a free user account at https://cropmanage.ucanr.edu/ before the workshop. Please arrive early to set up your laptop or tablet computer on the wifi and get logged on to CropManage.
Registration is free: Please register here
by April 1, 2024. Seats are limited to the first 30 registrants.
Questions: Contact Michael Cahn at mdcahn@ucanr.edu / 831-214-3690
Agenda
8:30 – 8:50 am Registration and computer set-up
8:50 – 9:20 am Introduction
9:20 – 10:00 am Getting started with CropManage
10:00 to 10:15 am break
10:10 – 11:00 am Using CropManage for decision support and record-keeping
11:00 – 11:45 am Group exercise
11:45- 12 pm Discussion /Q&A/ wrap up
Continuing Education Units (CEU) for Certified Crop Advisors (CCA) have been applied.
- Author: Michael D Cahn
Date: Thursday, March 7, 2024
Course 10:30 am – 2 pm
Drop-in office hours 2-3:30 pm
Location: Watsonville Public Library
275 Main St., Suite 100, Watsonville, CA 95076
- Learn how to use CropManage to support irrigation and nutrient management decisions and record-keeping for your crops.
- Learn how to set up your ranch and crops in CropManage.
- Learn about the latest updates and how CropManage can assist with Ag Order 4.0
- New—deep dive into advanced CropManage topics during office hours
CropManage is a free online decision-support tool for water and nutrient management of vegetables, berry, agronomic, and tree crops. Based on research and field studies conducted by the University of California Cooperative Extension, CropManage provides real-time recommendations for efficient and timely irrigation and fertilization applications while maintaining or improving overall yield.
At this free workshop, we will provide hands-on training so that you can learn to use the newest version of CropManage. Crops currently supported include many vegetables (carrots, cabbage, celery, broccoli, lettuce, tomato, spinach, etc.), berry crops (raspberry and strawberry), tree crops (almond, walnut, pistachio, prunes, and pear), and agronomic crops (alfalfa and corn). CropManage is also available in Spanish.
A new addition to the workshop are drop-in office hours for anyone already familiar with CropManage and does not want to attend the full program but needs more advance assistance or may have specific questions. Stop by at 2 pm and we will be ready to help answer your questions and dive deep into more advanced topics.
Who should participate? Growers, farm managers, other farm staff, crop advisors, consultants, and technical service providers are welcome. The workshop is for both new and current CropManage users. Spanish translation will be available. Lunch will be provided.
What to bring? This is a participatory workshop. Please bring a tablet or laptop computer so that you can follow along and participate in the exercises. Each participant will need a user account for CropManage. Please set up a free user account at https://cropmanage.ucanr.edu/ before the workshop. Please arrive early to set up your laptop or tablet computer on the wifi and get logged on to CropManage.
Registration is free: Please register here
by March 4, 2024. Seats are limited to the first 30 registrants.
Questions: Contact Michael Cahn at mdcahn@ucanr.edu / 831-214-3690 or Sacha Lozano at slozano@rcdsantacruz.org / 831-224-0293
Agenda
10:30 – 10:50 am Registration and computer set-up
10:50 – 11:20 am Introduction
11:20 – 12:00 pm Getting started with CropManage
12:00 to 12:30 pm Lunch break
12:30 – 1:15 pm Using CropManage for decision support and record-keeping
1:15 – 1:45 pm Group exercise
1:45- 2 pm Discussion /Q&A/ wrap up
Drop-in office hours
2 – 3:30 pm Stop by to receive one-on-one assistance with CropManage and explore more advanced ways to use CropManage to understand nutrient and water needs of crops.
Continuing Education Units (CEU) for Certified Crop Advisors (CCA) have been applied.